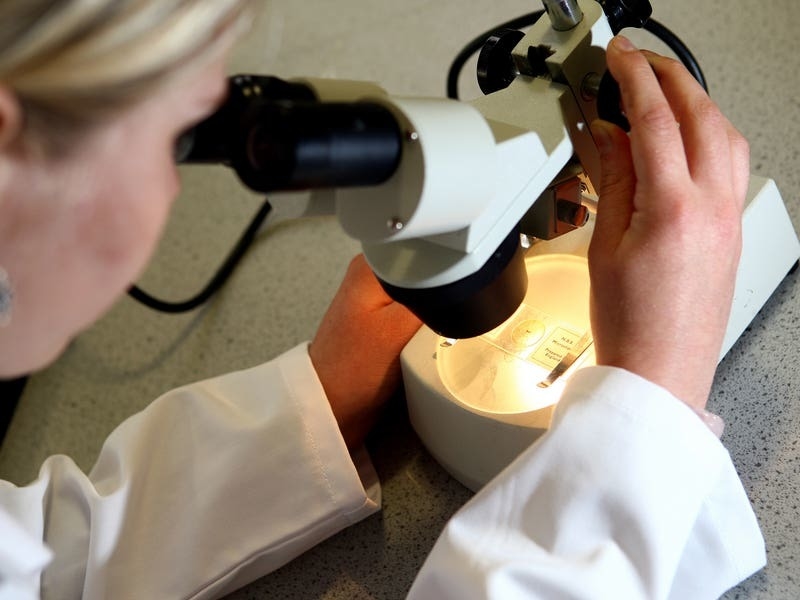
Scientists have released a new “pangenome” sequence in a bid to try and understand more about the differences between people’s genes.
A pangenome combines the genome sequences of many people which means that it is easier to see all of the genetic diversity and differences in that collection.
Researchers from the international Human Pangenome Reference Consortium have announced that they have mapped out the genomes of 47 people in the new pangenome sequence, and aim to increase this number to 350 by the middle of next year.
Experts hope that the work, which has been published as a collection of papers in the journal Nature, will lead to a “a new age of genetic diagnosis”.
Researchers from the @genome_gov-funded Human Pangenome Reference Consortium (@HumanPangenome) have completed a collection of new human reference genome sequences that much more accurately reflect global diversity! https://t.co/jaercozEem pic.twitter.com/ZkjSJH8NAE
— National Human Genome Research Institute (@genome_gov) May 10, 2023
When researchers and clinicians study people’s DNA, their genetic information is compared to a “reference” sequence of genomes.
The human reference genome was first completed 20 years ago.
Despite many revisions to the sequence, the three billion letters of DNA code that make up this reference genome were put together using information from just 20 people who lived in the same part of North America, and most of the reference sequence is from only one person.
Any two people’s genomes are, on average, more than 99% identical.
But the small differences are what make people different from one another and can provide insights about their health including how to predict outcomes, diagnose disease and guide medical treatments.
“Everyone has a unique genome, so using a single reference genome sequence for every person can lead to inequities in genomic analyses,” said co-author of the main study, Adam Phillippy, from the National Human Genome Research Institute in the US.
“For example, predicting a genetic disease might not work as well for someone whose genome is more different from the reference genome.”
The new pangenome sequence builds upon the previous reference genome sequence, adding more than 100 million new bases, or “letters” in DNA, academics said.
The previous reference genome is described as “linear” while the new pangenome represents many different versions of the human genome sequence at the same time and has been likened to a “map of a subway system”.
Eric Green, director of the National Human Genome Research Institute, added: “Basic researchers and clinicians who use genomics need access to a reference sequence that reflects the remarkable diversity of the human population. This will help make the reference useful for all people, thereby helping to reduce the chances of propagating health disparities.”
Commenting, Professor David Adelson, from the University of Adelaide in Australia, said: “Why is a pangenome so exciting? It includes all the differences between the genomes of the individuals that have been sequenced.
“All you have to do is look around you to see how different people are. These differences reflect differences in our genomes. Up until now we have used a single genome sequence as a reference for the detection of genetic changes that cause disease. That reference did not include differences between people or populations.
“With the pangenome we can now look for genetic changes across many individuals and ultimately the pangenome will grow to include information from thousands and perhaps millions of genome sequences.
“This means our ability to use genetic information for diagnosis will increase enormously. With the current pangenome from only 45 humans, the accuracy of detection to find genetic changes has gone up by 34% and the number of large, difficult-to-detect changes we now know about has gone up by over 100%.
“This paper heralds a new age of genetic diagnosis, that will benefit people from all ancestries, unlike our current reference genome that does not reflect all the diversity of humanity.”